この節の作者: Rebecca Vederhus, Sebastian Jentschke
From SPSS to jamovi: Analysis of frequencies¶
This comparison shows how a loglinear analysis is conducted in SPSS and jamovi. The SPSS test follows the description in chapter 19.9.2 in Field (2017), especially figure 19.7 and output 19.7 - 19.10. It uses the data set Cats and Dogs.sav which can be downloaded from the web page accompanying the book.
| SPSS | jamovi |
|---|---|
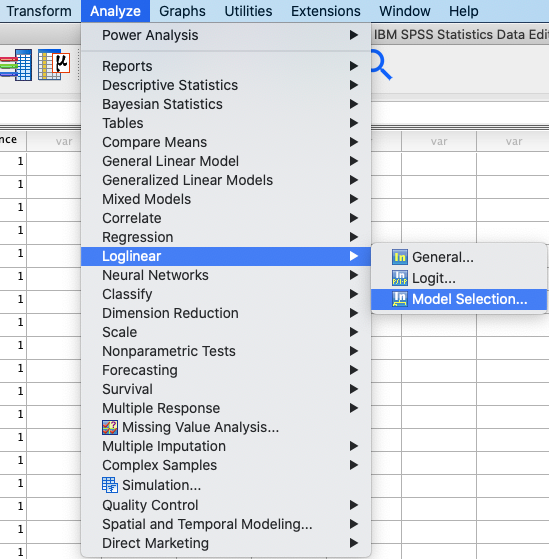
In SPSS, you can run a loglinear analysis using: Analyze → Loglinear
→ Model Selection. |
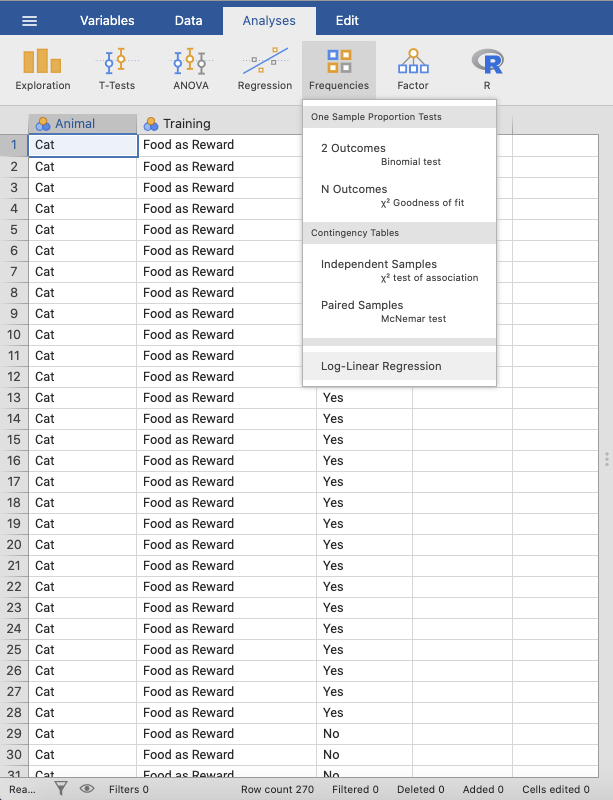
In jamovi, this can be done using: Analyses → Frequencies → Log-
Linear Regression. |
 |
 |
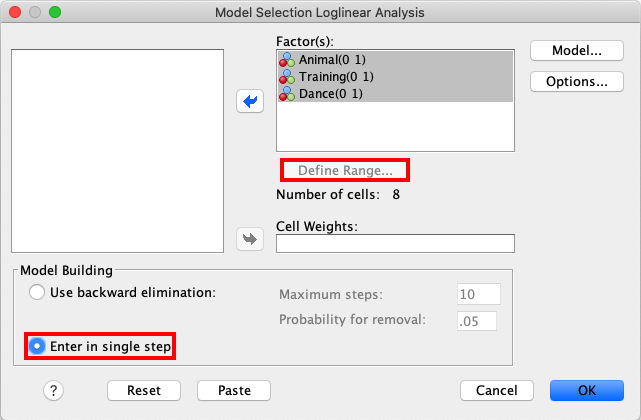
In SPSS, move the variables Animal, Training and Dance to the
Factor(s) box. Then, mark all three variables and click Define Range.
In this window, set Minimum as 0 and Maximum as 1. Click
Continue. In the box called Model Building, click Enter in single
step. |
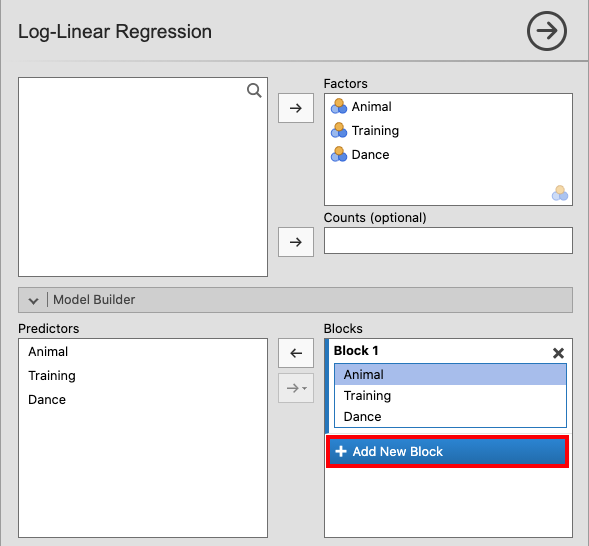
In jamovi, move Animal, Training and Dance to Factors. Open
the Model Builder window, click + Add New Block and move the three
variables to this block. |
 |
 |
 |
|
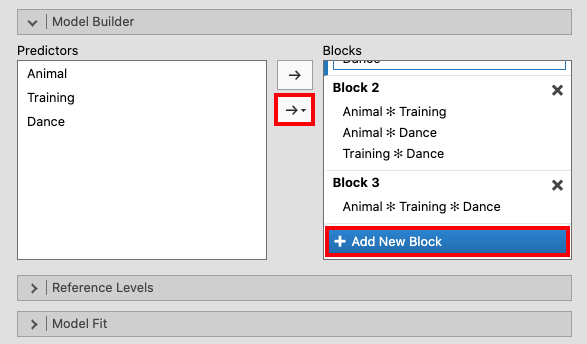
Add another block. Mark all three variables and choose All 2 way from the
drop-down menu. Then, add a third block and mark all three variables. Open
the drop-down menu and click All 3 way. |
|
 |
|
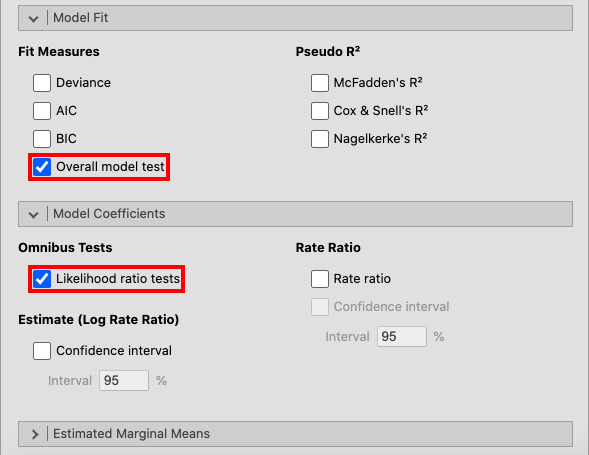
Open Model Fit and tick the box for Overall model test. Lastly, tick
Likelihood ratio tests in the Model Coefficients window. |
|
 |
|
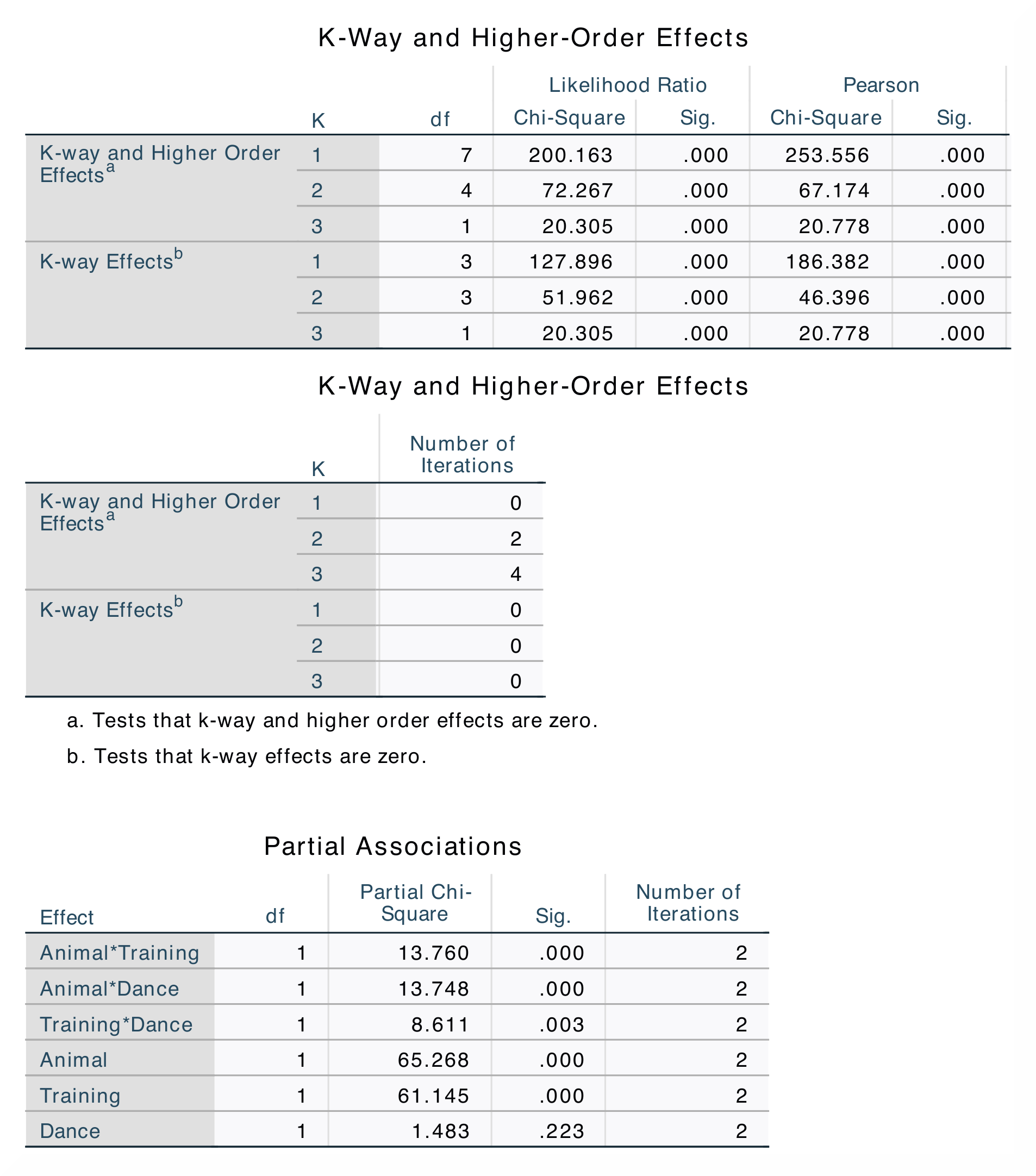
Only the results in the output tables K-Way and Higher-Order Effects and Partial Associations in SPSS are replicated in the jamovi analysis. |
|
 |
 |
 |
|
In the K-Way and Higher-Order Effects table, you can find df-values,
likelihood ratio statistics and significance values when K = 1, 2 and 3. The
SPSS results also contains Pearson chi-square statistics. The different rows
show if any higher-order effects or one-way effects significantly affect the
model fit. The Partial Associations table breaks the model into specific
parts and tells us which two-way interactions that significantly affect the
fit of the model. You can tell this by looking at the significance values
for the different interactions and comparing them. |
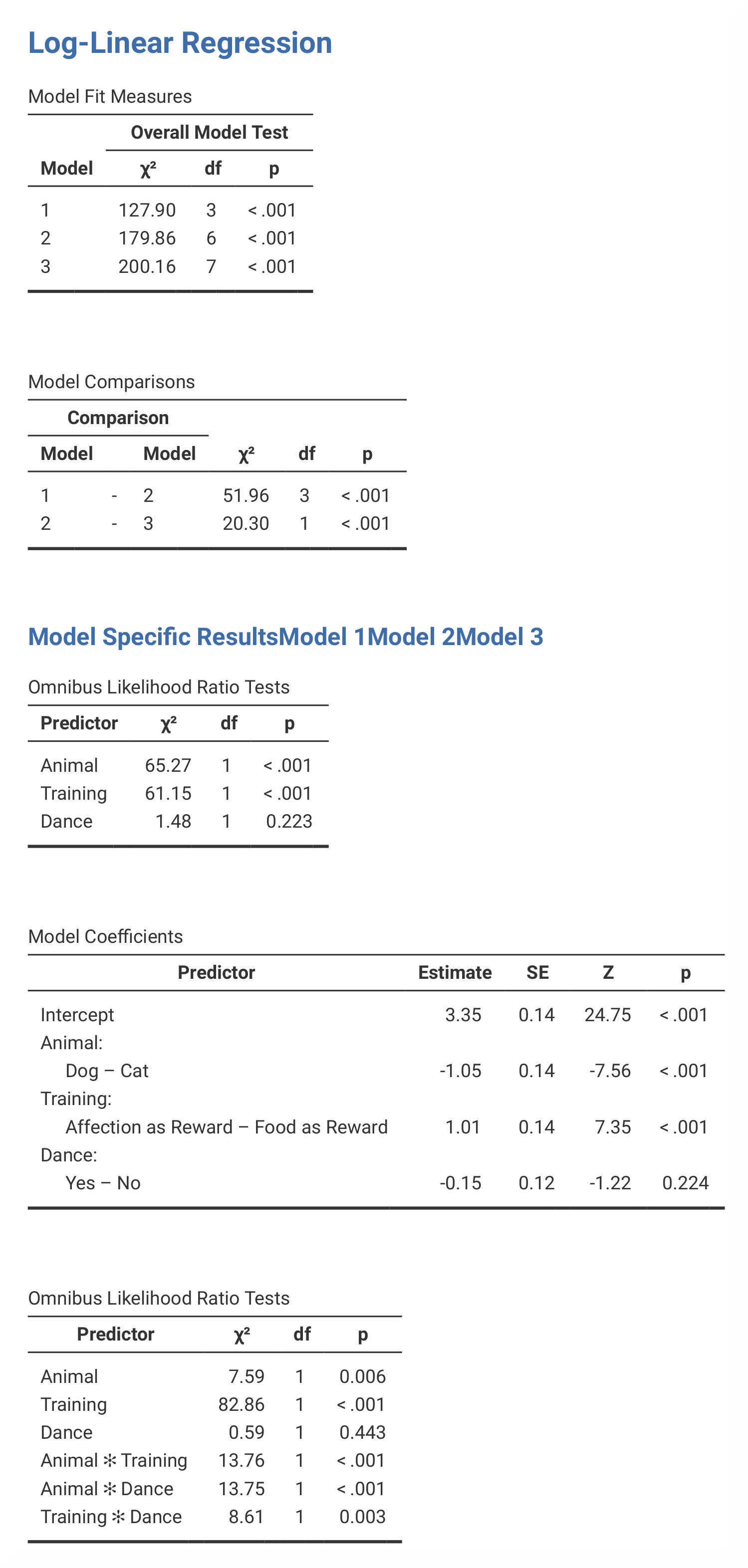
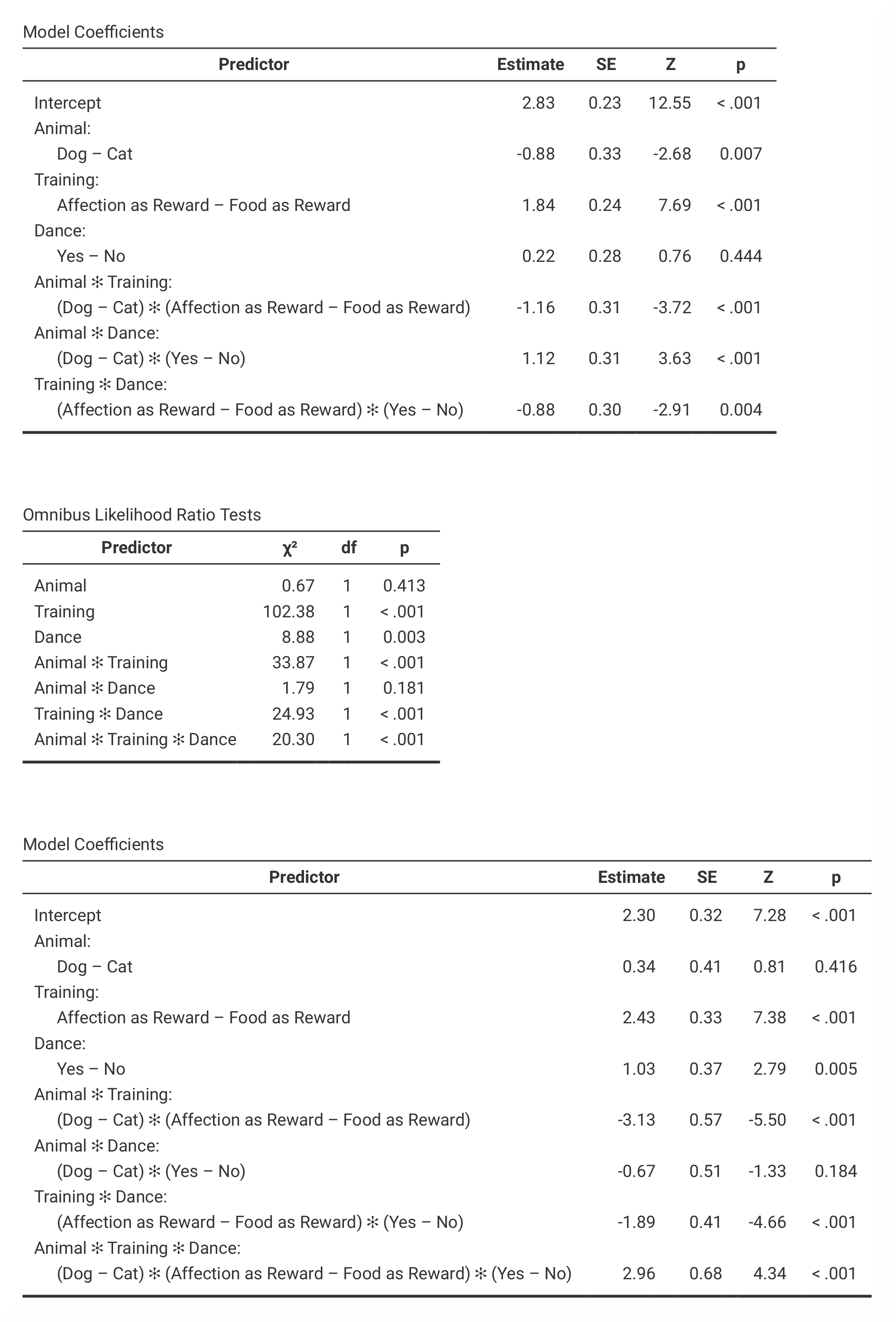
In jamovi, the values that are found in the K-Way and Higher-Order
Effects table in SPSS can be found in the Model Fit Measures and
Model Comparisons tables. However, jamovi does not provide Pearson
chi-square statistics and the number of iterations. The partial associations
table in jamovi is called Omnibus Likelihood Ratio Tests and are
presented in three separate tables (one for each model). |
Output from the SPSS analysis contains a lot of tables that are not included in the jamovi analysis. In addition, the results from the parameter estimates tables differ from each other, and are therefore not included here. The numerical values for the statistics are the same: χ² = 127.90, p < .001; χ² = 200.16, p < .001; χ² = 51.96, p < .001: χ² = 20.30, p < .001; χ² = 65.27, p < .001; χ² = 61.15, p < .001; χ² = 1.48; χ² = 13.76, p < .001; χ² = 13.75, p < .001; χ² = 8.61, p < .01. |
|
| If you wish to replicate those analyses using syntax, you can use the commands below (in jamovi, just copy to code below to Rj). Alternatively, you can download the SPSS output files and the jamovi files with the analyses from below the syntax. | |
HILOGLINEAR Animal(0 1) Training(0 1) Dance(0 1)
/CRITERIA ITERATION(20) DELTA(.0)
/PRINT=FREQ RESID ASSOCIATION ESTIM
/DESIGN.
|
jmv::logLinear(
data = data,
factors = vars(Animal, Training, Dance),
blocks = list(
list("Animal", "Training", "Dance"),
list(c("Animal", "Training"), c("Animal", "Dance"),
c("Training", "Dance")),
list(c("Animal", "Training", "Dance"))),
refLevels = list(
list(var = "Animal", ref = "Cat"),
list(var = "Training", ref = "Food as Reward"),
list(var = "Dance", ref = "No")),
modelTest = TRUE,
dev = FALSE,
aic = FALSE,
pseudoR2 = NULL,
omni = TRUE)
|
| SPSS output file containing the analyses | jamovi file containing the analyses |
References
Field, A. (2017). Discovering statistics using IBM SPSS statistics (5th ed.). SAGE Publications. https://edge.sagepub.com/field5e