この節の作者: Sebastian Jentschke, Jonathon Love
The jmv R package¶
For some use-cases, it's preferable to be able use the jamovi analyses from R. This is provided by thejmvR package. All the analyses included with jamovi are available from within R using this package.
Preparation¶
jmvis available from CRAN (a central repository for publicly available R-packages). To install jmv, open an interactive R session (e.g. RStudio) and type the first line (the >-mark at the begin of the line is the input-mark of R and must not be copied / typed). The packagehavenis required if you want to read or write files from SPSS, SAS or Stata and can be installed using the second line. If you want to read and write jamovi's.omv-files you can use the packagejmvReadWritewhich is installed using the third line. Finally, the packagejmvconnectpermits to access data sets that you currently have opened in jamovi.More information about the analyses
jmvprovides or how to use the other packages is available using the R help system, or from the overview below.install.packages('jmv') install.packages('haven') install.packages('jmvReadWrite') install.packages('jmvconnect')
Use of jamovi syntax in R¶
For examples on how to use
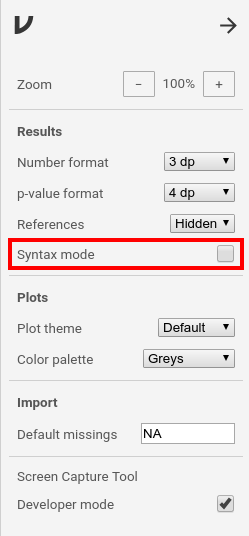
jmv, jamovi can be placed in ‘Syntax mode’. Syntax mode produces the R syntax required to reproduce jamovi analyses in R. To enable the syntax mode, you have to click the properties icon (⋮) in the top-right corner. Set a tick at syntax mode in the properties window. Close the properties with the arrow-icon (→) at the top-right.
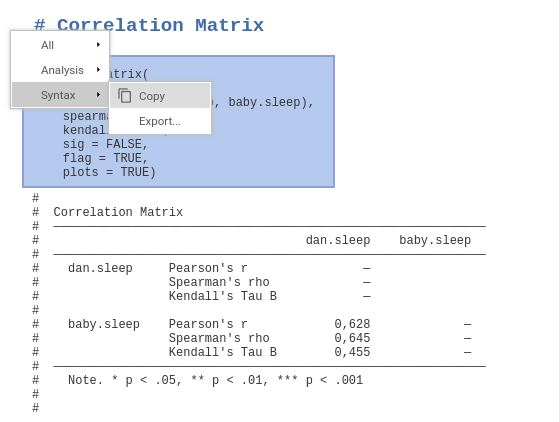
The main window changes to text mode and you can run analyses and afterwards right-click on the command the appears at the top of each analysis to export or copy the syntax.
Alternatively, you can write syntax directly. To do this, open R or RStudio and type the command in the first line. As during the description of the installation above; the second line is required if you want to use files from SPSS, SAS or Stata; the third line for reading jamovi's
.omv-files, and the fourth line for directly accessing datasets opened in your current jamovi-session.library(jmv) library(haven) library(jmvReadWrite) library(jmvconnect)Afterwards, you are ready to analyze your data. Typically, you have to load a dataset first. Use the first line if you have a
.csv-file (sephas to be set to the separator between data cells, e.g.,,,;, etc.), the second line for loading SPSS'.sav-files, or the third line for loading jamovi's.omv-files:data = read.csv("data.csv", header = TRUE, sep = ",") data = haven::read_sav("data.sav") data = jmvReadWrite::read_omv("data.omv")With those data, you can run whatever analysis you like (you will find an overview of available functions a little further down on this page). For example, to run a simple descriptive-statistics-analysis:
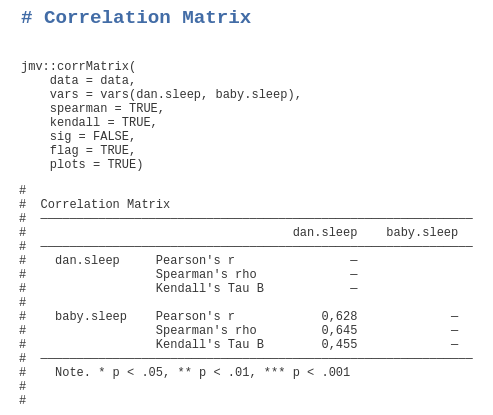
jmv::descriptives(data = data, vars = vars(var1, var2))or for a correlation between to variables (quite basic in the first and more advanced - adding two non-parametric measures and plots - in the second line; please note that pearson = TRUE is not necessary because it is the default):
jmv::corrMatrix(data = data, vars = vars(var1, var2), pearson = TRUE, sig = TRUE) jmv::corrMatrix(data = data, vars = vars(var1, var2), spearman = TRUE, kendall = TRUE, sig = FALSE, flag = TRUE, plots = TRUE)